What is Strandedness in RNA-seq?
What is RNA strandedness?
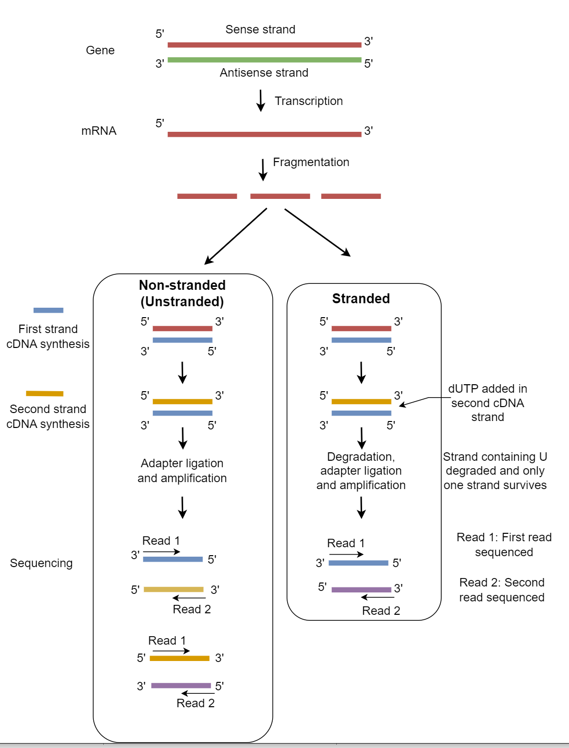
Based on the RNA sequencing (RNA-seq) library preparation protocol, the strandedness in RNA-seq can be classified as stranded (strand-specific) RNA-seq and non-stranded (Unstranded) RNA-seq.
Stranded RNA-seq preserves the orientation (direction) of original mRNA transcripts, making it possible to identify whether a sequence read originates from the sense strand or the antisense strand.
The non-stranded RNA-seq method, however, does not preserve the orientation of the original mRNA transcripts. This means you will not be able to determine whether the sequence read originates from the sense or antisense strand.
Stranded RNA-seq library preparation is similar to non-stranded RNA-seq, except that it includes additional steps such as adding dUTPs during second-strand cDNA synthesis and digesting uracil-containing strands (U).
Follow this article, if you would like to determine the strandedness in your RNA-seq data.
Advantages of Stranded RNA-seq
Stranded RNA-seq is a powerful method to study transcriptome profiling and it has several advantages including,
- Accurately quantifying gene expression levels for overlapping genes that are transcribed from opposite strands. This means you can determine whether the read is coming from sense or antisense transcripts.
- Identify novel antisense transcripts and precisely annotate the genome
- Identify gene fusions and non-coding RNAs
- Allele-specific expression analysis
- Higher gene assembly
Enhance your skills with courses on genomics and bioinformatics
- Genomic Data Science Specialization
- Biology Meets Programming: Bioinformatics for Beginners
- Python for Genomic Data Science
- Bioinformatics Specialization
- Command Line Tools for Genomic Data Science
- Introduction to Genomic Technologies
This work is licensed under a Creative Commons Attribution 4.0 International License
Some of the links on this page may be affiliate links, which means we may get an affiliate commission on a valid purchase. The retailer will pay the commission at no additional cost to you.

